Memory Stack
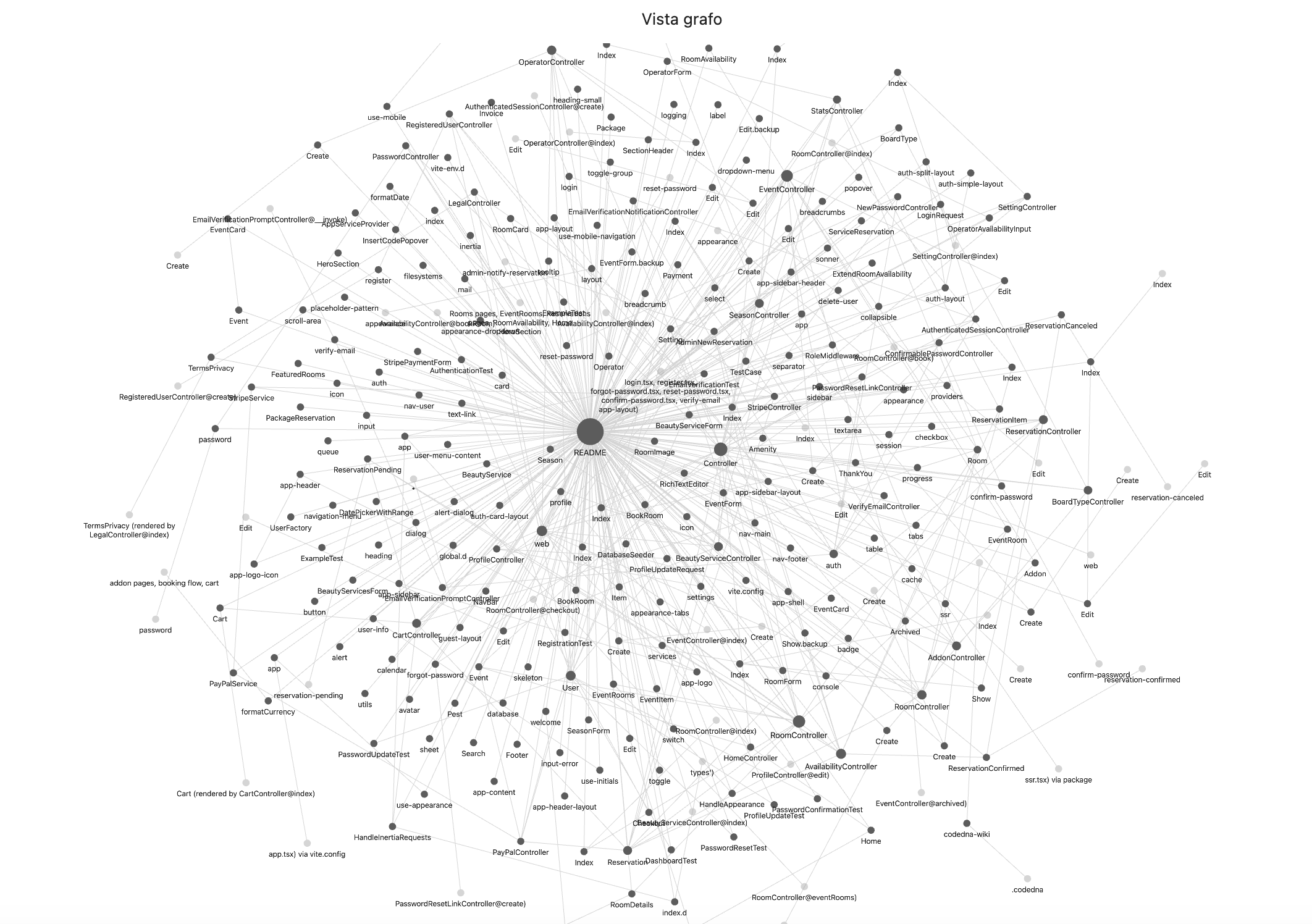
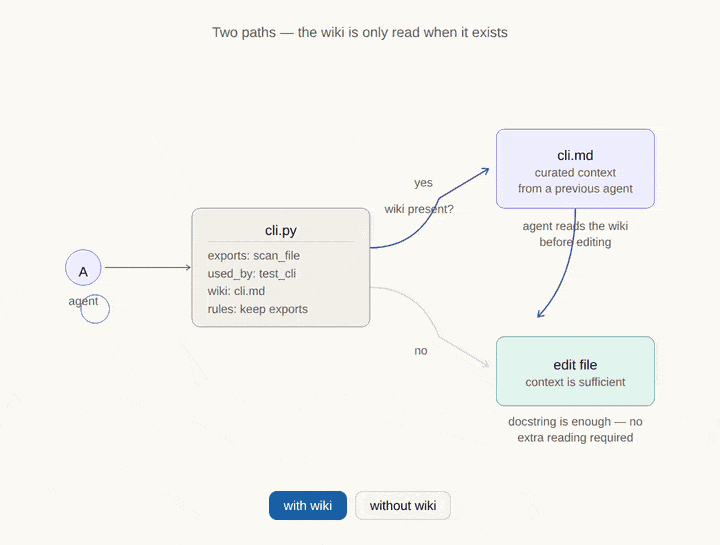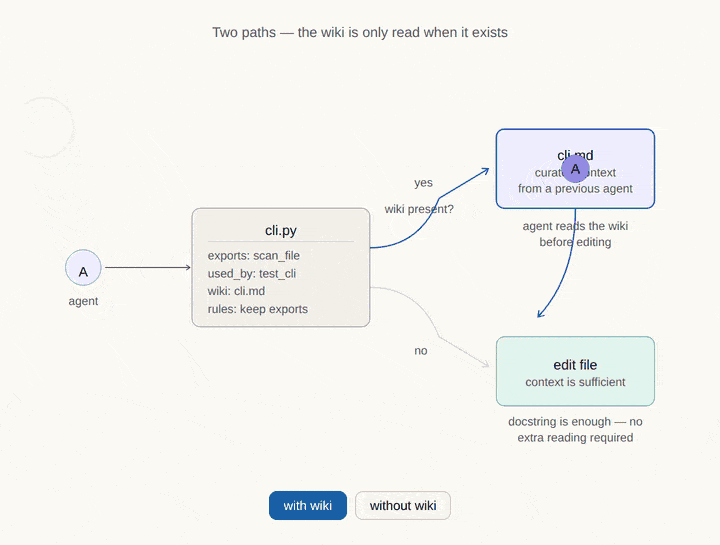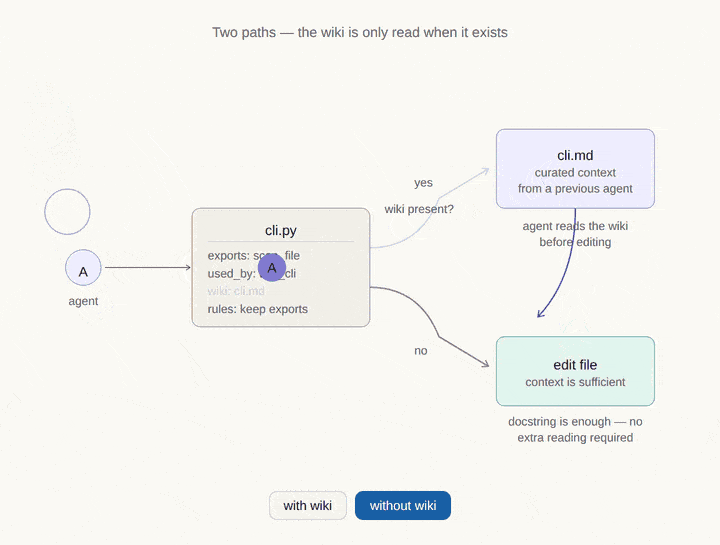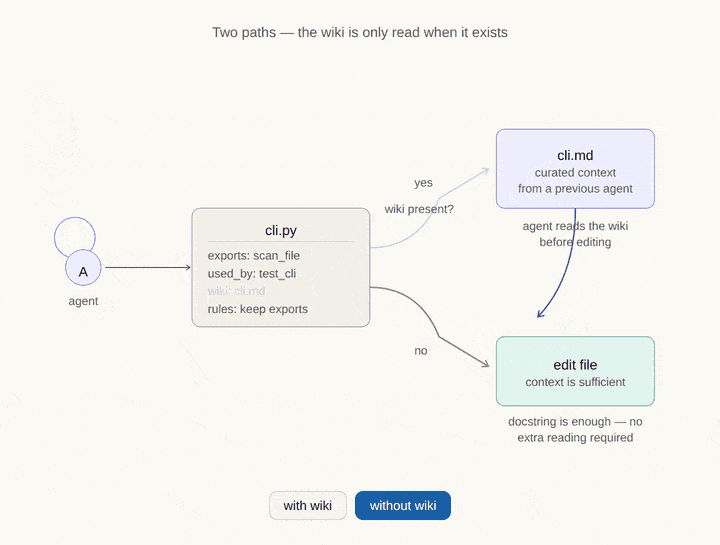
Where CodeDNA sits in the AI memory stack.
Every AI coding agent relies on multiple memory layers. Most of them are external to the code. CodeDNA is the only layer that lives inside the source files themselves — it travels with every clone, fork, and CI pipeline.
| Layer |
Examples |
Where it lives |
Shared across tools? |
| LLM / Agent |
Claude, GPT-4, Cursor, Copilot |
Cloud |
— |
| External memory |
Chat history, Memory API |
Cloud / external DB |
✗ tool-specific |
| Markdown / Config |
CLAUDE.md, .cursorrules, AGENTS.md |
Repo (outside source files) |
partial |
| CodeDNA |
exports, rules, agent, message, .codedna |
Inside every source file + repo root |
always |
Multi-Model SWE-bench
Tested across multiple LLMs.
Django issues from SWE-bench, tested across multiple LLMs. Same prompt, same tools, same tasks. DeepSeek Chat: +17pp F1, p=0.001, 10/0/0 · Gemini 2.5 Flash: +13pp F1, p=0.040 · Gemini 2.5 Pro: +9pp F1. All 3 models improve.
File Localization F1 — Control vs CodeDNA by Model
Navigation Demo — django__django-11808 · DeepSeek Chat · 5 runs
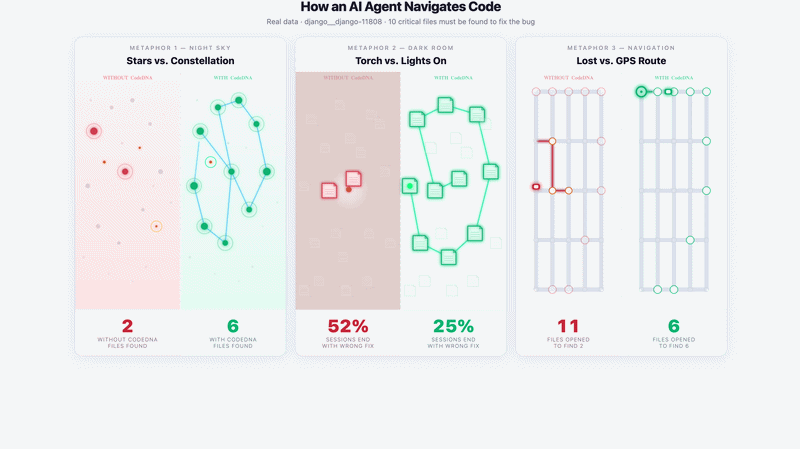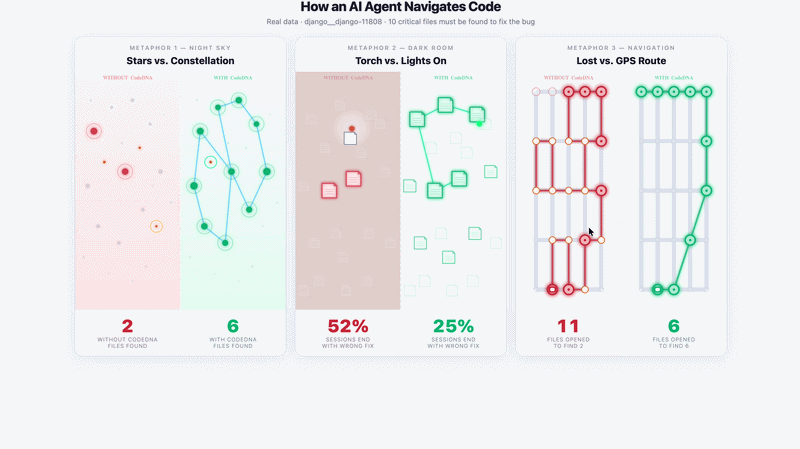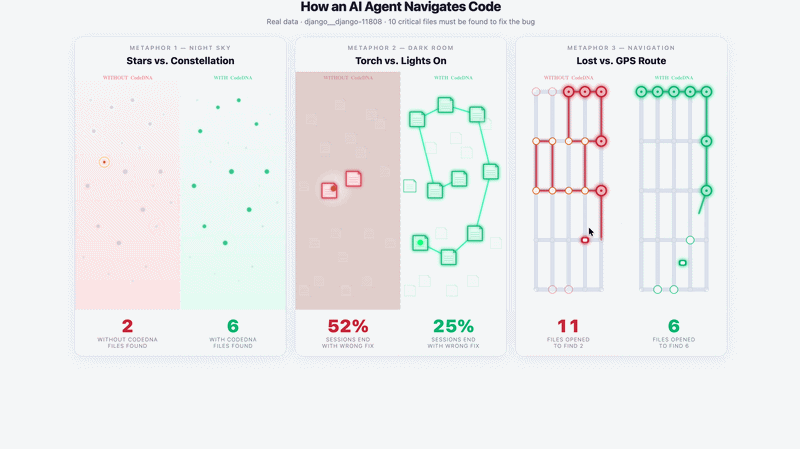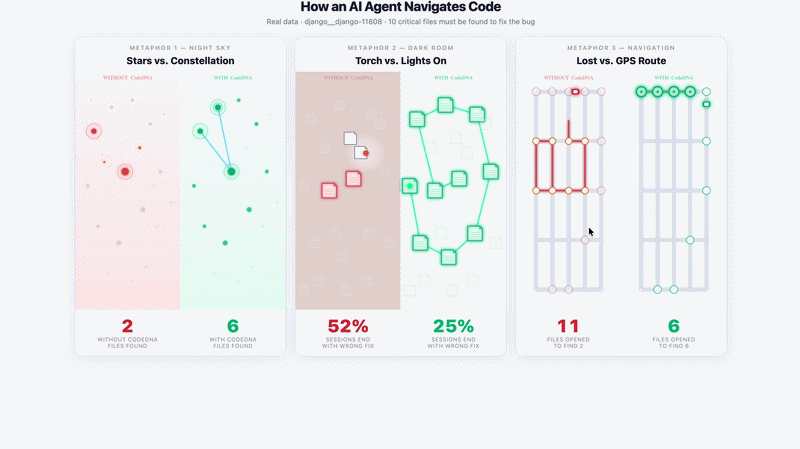
Without CodeDNA: agent opens random files, stops early — 2/10 critical files found. |
With CodeDNA: follows used_by: chain — 6/10 critical files found.
Best Model
🏆 Gemini 2.5 Flash — +13pp F1
From 60% to 72%. Wins 4 out of 5 tasks. Δ up to +21pp on delegation chains (Task 13495). p=0.040, Wilcoxon W+=14, N=5 tasks × ≥5 runs at T=0.1.
DeepSeek Chat
DeepSeek Chat — +17pp F1 (p=0.001 · 10/0/0)
From 50% to 60%. Wins 4/5 tasks. Notable: +35pp on cross-cutting task 11808 — opposite direction from Gemini Flash (−1pp). Task 13495 anomaly (−9pp) under investigation. Not statistically significant.
Key Finding
🧠 Model-Agnostic Benefits
4 out of 5 models improve with CodeDNA. The benefit is strongest on tasks requiring cross-module navigation — exactly where AI agents struggle most.
🔬 Methodology: SWE-bench Django tasks × 3 models (Gemini 2.5 Flash ✓, DeepSeek Chat 10 tasks ✓, Gemini 2.5 Pro ✓). 3–5 runs/task at T=0.1. Identical system prompt, same 3 tools (read_file, list_files, grep), max 30 turns. Metric: File Localization F1 (ground-truth files from patch). Statistical test: Wilcoxon signed-rank (one-tailed). 6 DeepSeek tasks independently replicated by @fabioscialanga. Script: benchmark_agent/swebench/run_agent_multi.py.